This web page was produced as an assignment for Gen677 at UW-Madison Spring 2012.
Protein Interaction Networks
Protein-protein interaction networks were obtained from STRING 9.0 which is a database of known and predicted protein-protein interactions. Below are the protein-protein interaction networks for human and mouse ITGAM. The colored lines indicate that the proteins have been found to interact in research studies. The light gray lines indicate that the proteins have a connection based on primary literature abstracts but have not yet been shown to physically interact in research studies.
Human ITGAM protein-protein interaction network
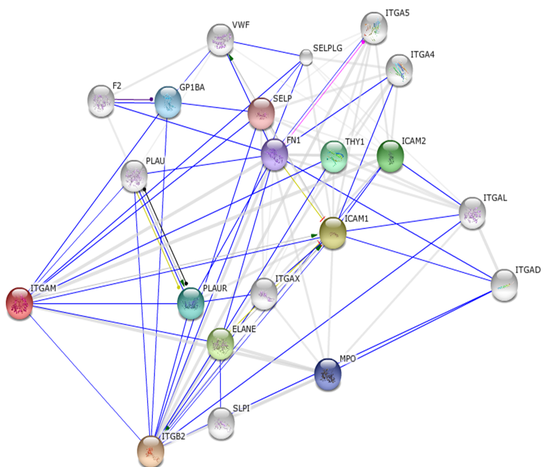
Mouse ITGAM protein-protein interaction network
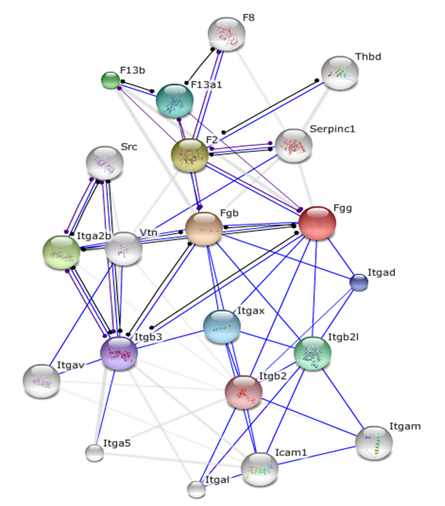
Analysis
Many proteins found in the STRING protein-protein interaction networks overlap between mouse and human ITGAM. The similarity between the two ITGAM networks provides support for the use of the SLE murine model when researching the role of ITGAM in the pathogenesis of SLE.
Interestingly, the mouse ITGAM interaction network contains F2, FGG, and Src. These are some of the proteins I focus on in my Conclusion and Future Directions. These proteins are the focus of my proposed model for how the ITGAM, estrogen, and coronary heart disease pathways are intertwined. This is additional support for the use of the murine model in the investigation of the role of ITGAM in the pathogenesis of SLE.
The differences in the two networks may have to do with slight variations in the ITGAM pathway and/or lack of scientific research.
Resources
STRING 9.0 http://string-db.org/
Created by: Stephanie Kroll, [email protected], Page last updated: 5/22/12, Genetics 677 Class Web Page
