This web page was produced as an assignment for Gen677 at UW-Madison Spring 2012.
Phylogeny
A phylogenetic tree depicts lineages of organisms over time. The following trees were made by comparing the similarity between the protein sequences of ITGAM homologs. Branch points represent a common ancestor of the two diverging species.
Phylogenetic trees for ITGAM protein homologs
MUSCLE Phylogeny
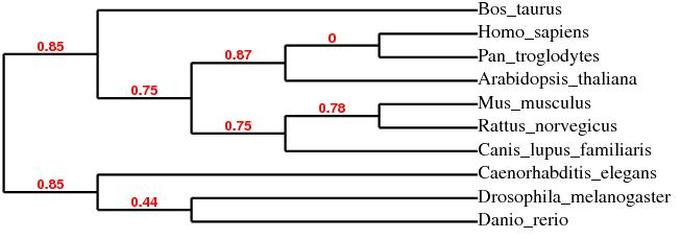
ClustalW2 Phylogeny
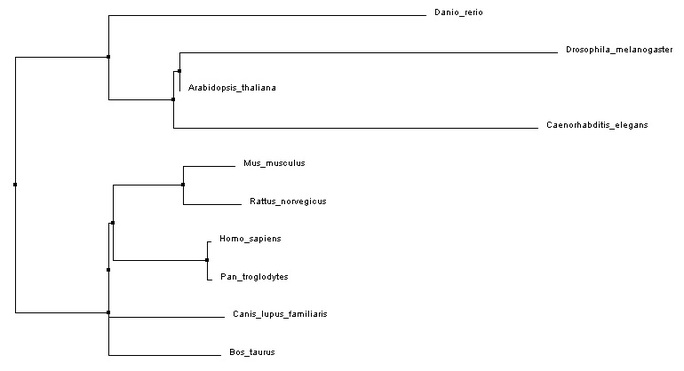
Analysis
The MUSCLE and ClustalW2 phylogenetic trees do not agree completely. However, both trees agree that the ITGAM chimpanzee homolog is the closest relative to human ITGAM of all the protein homologs analyzed. Opposite of what would be expected based on the the protein domain similarity of the homologs, not all of the vertebrate homologs cluster together in the phylogenetic trees above. The majority of vertebrate homologs cluster together, but Danio rerio (zebra fish) does not cluster with the other vertebrate homologs. Even though the Danio rerio homolog contains the same domains as human ITGAM, it does not have as much sequence similarity as the other vertebrate homologs.
References
Created by: Stephanie Kroll, [email protected], Page last updated: 5/22/12, Genetics 677 Class Web Page
