This web page was produced as an assignment for Gen677 at UW-Madison Spring 2012.
Microarray experiments
Microarrays are used to measure expression levels (how much a particular is transcribed into mRNA from DNA) of multiple genes at one time using DNA probes linked to fluorophores. The following microarray experiments analyze the expression of ITGAM, FGG, or STAT3 by determining the levels of mRNA under various estrogen conditions. These data sets used Affymetrix chips and were found on GEO Profiles. The expression levels are measured in arbitrary units.
GEO microarray profile for ITGAM expression in human breast cancer MCF-7/BUS cells treated with 17beta-estradiol (E2) at concentrations up to 100 pM
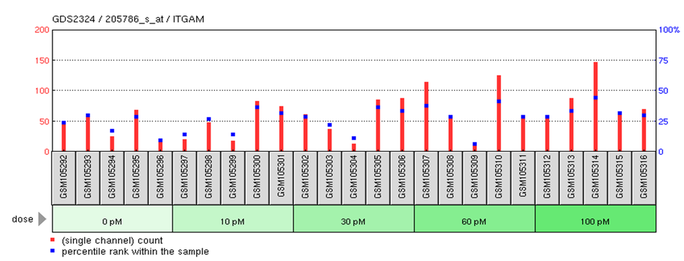
GEO microarray profile for FGG expression in human MCF-7 breast cancer cells treated with estrogen for up to 12 hours
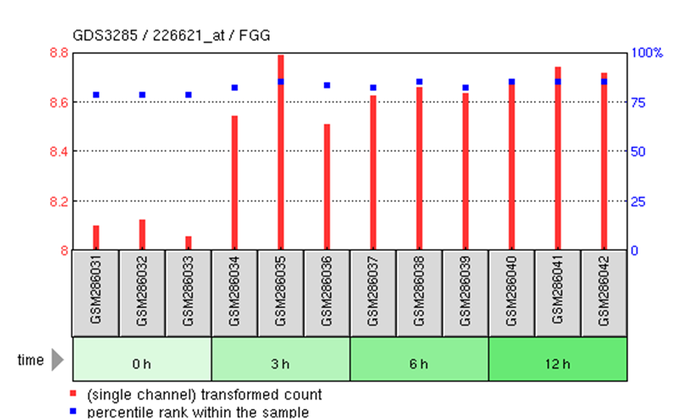
GEO microarray profile for STAT3 in MCF-7 human breast cancer cells treated with estrogen up to 12 hours
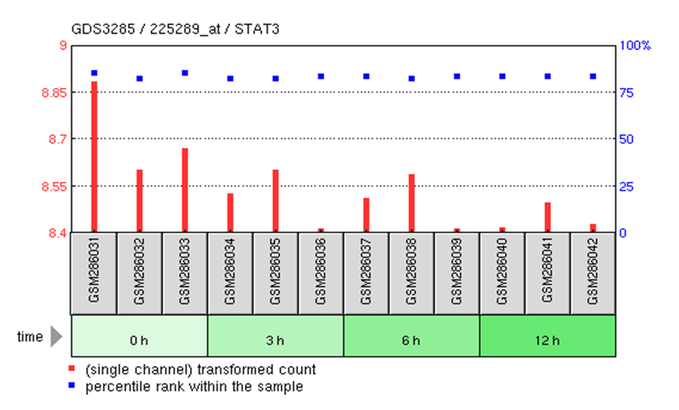
Analysis
After finding proteins that could be the links between ITGAM, SLE, estrogen, and an increased risk for coronary heart disease in patients with lupus (described on Interaction networks), I decided to find microarray data on GEO Profiles for these proteins. I was only able to find microarray data on ITGAM, FGG, and STAT3 expression in the presence and absence of estrogen in human breast cancer cells (MCF-7). The microarray data for these experiments are presented above.
Summary of the microarray data above:
- Estrogen does not appear to alter ITGAM expression.
- Estrogen appears to increase FGG expression.
- Estrogen appears to decrease STAT3 expression.
After comparing this data to the Interaction networks, it appears as though STAT3 could be a negative regulator of FGG expression. A proposed model for how these proteins and the other proteins found in the Interaction networks is on the Conclusion and Future Directions page.
References
GEO Profiles http://www.ncbi.nlm.nih.gov/geo/
Created by: Stephanie Kroll, [email protected], Page last updated: 5/22/12, Genetics 677 Class Web Page
